Efficiently analyze and maximize the potential of your immune repertoire sequencing data.
ImmunoRaptor – Immune Repertoire Analysis Software
Over the past few years, it has become easier to study immune responses through adaptive immune receptor repertoire sequencing (AIRR-Seq), in which DNA from B-cell or T-cell receptors is sequenced. However, there is a lack of easy-to-use, efficient, and accurate tools for analyzing these data.
Therefore, Excelra has developed ImmunoRaptor, a highly innovative immune repertoire analysis software that enables researchers to easily manage and analyze their immune repertoire sequencing data. Its wide range of features ensures that ImmunoRaptor can be applied in diverse domains such as antibody discovery, autoimmune diseases, vaccine development, and oncology. ImmunoRaptor has an easy-to-use interface and does not require bioinformatics expertise. By relying on cloud computing, hundreds of samples can be quickly analyzed simultaneously.
Would you like to try out ImmunoRaptor for your immune repertoire sequencing data?
6 easy steps to fully benefit from your immune repertoire sequencing data
Benefits of ImmunoRaptor
No bioinformatics expertise required
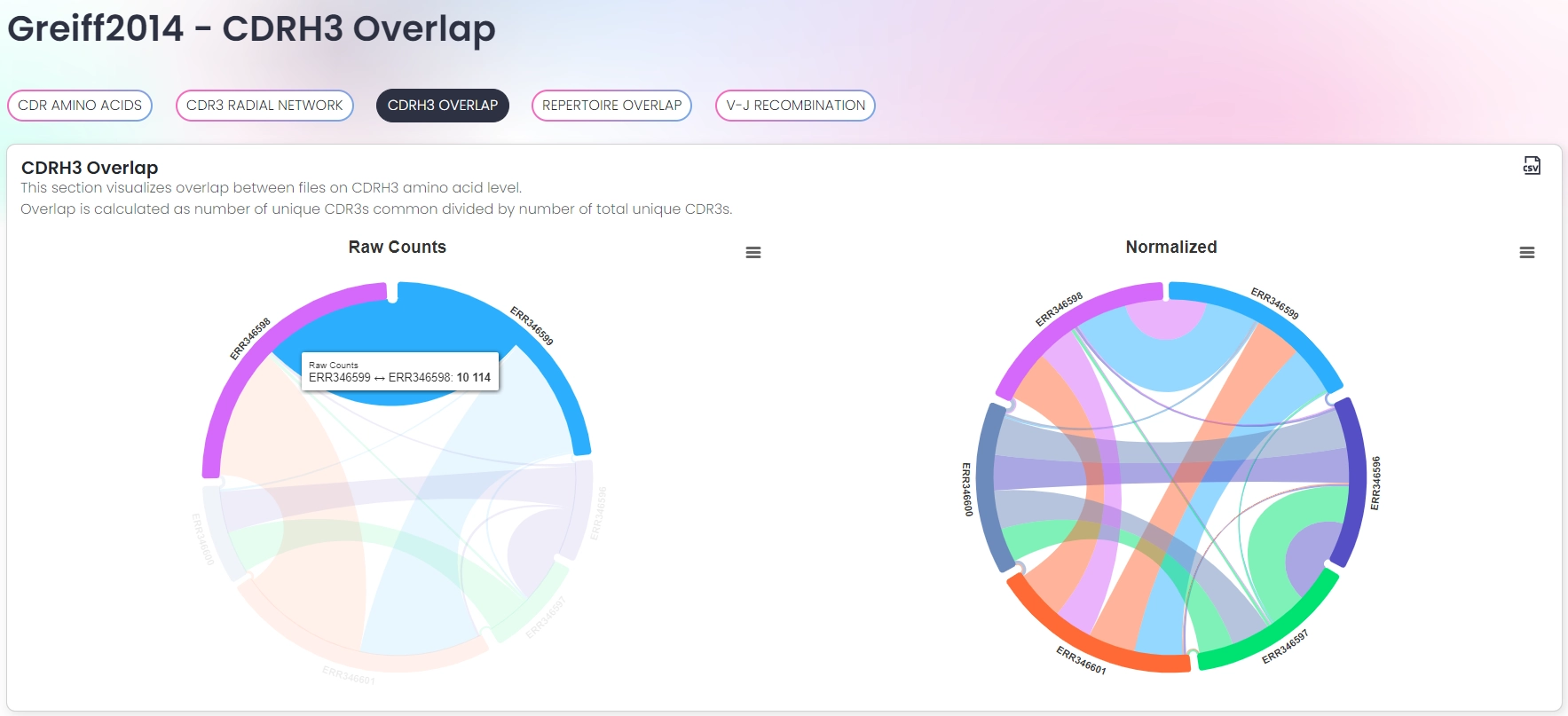
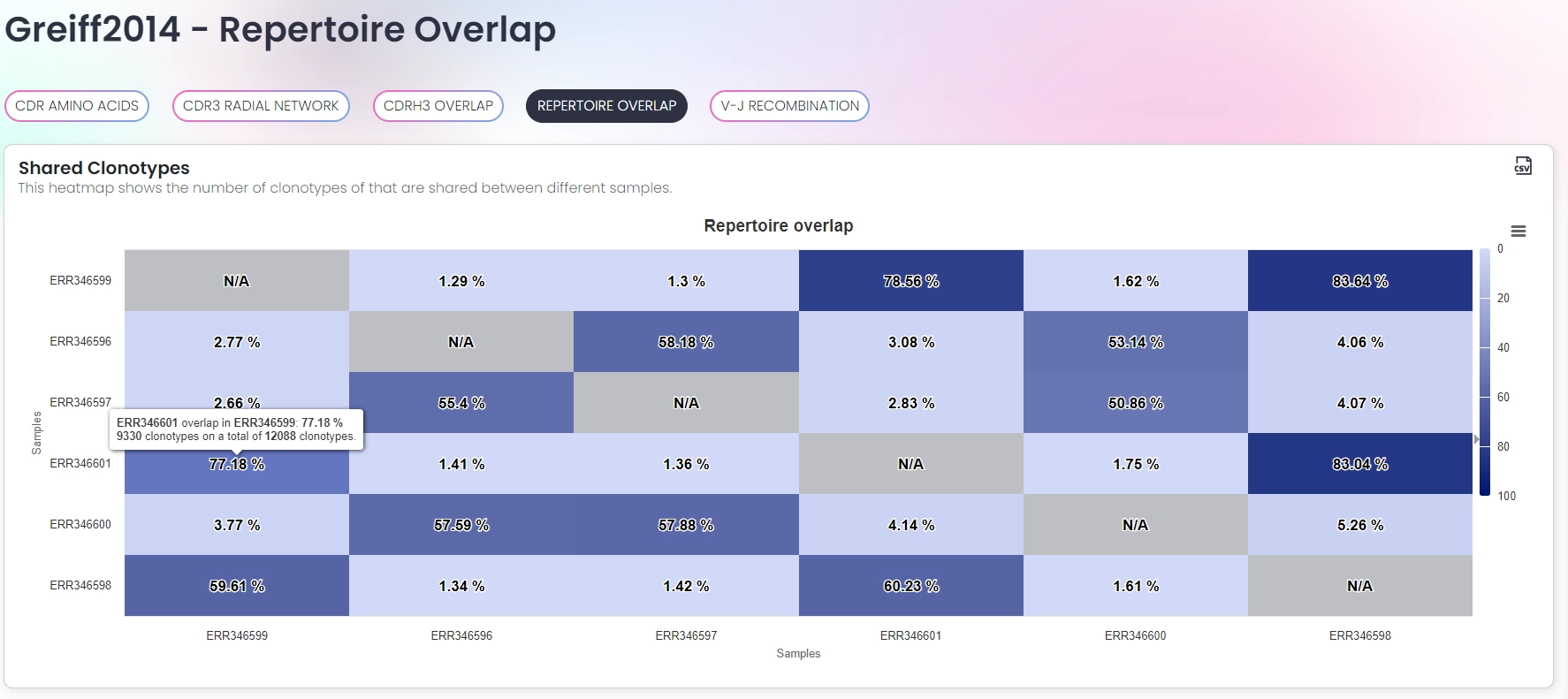
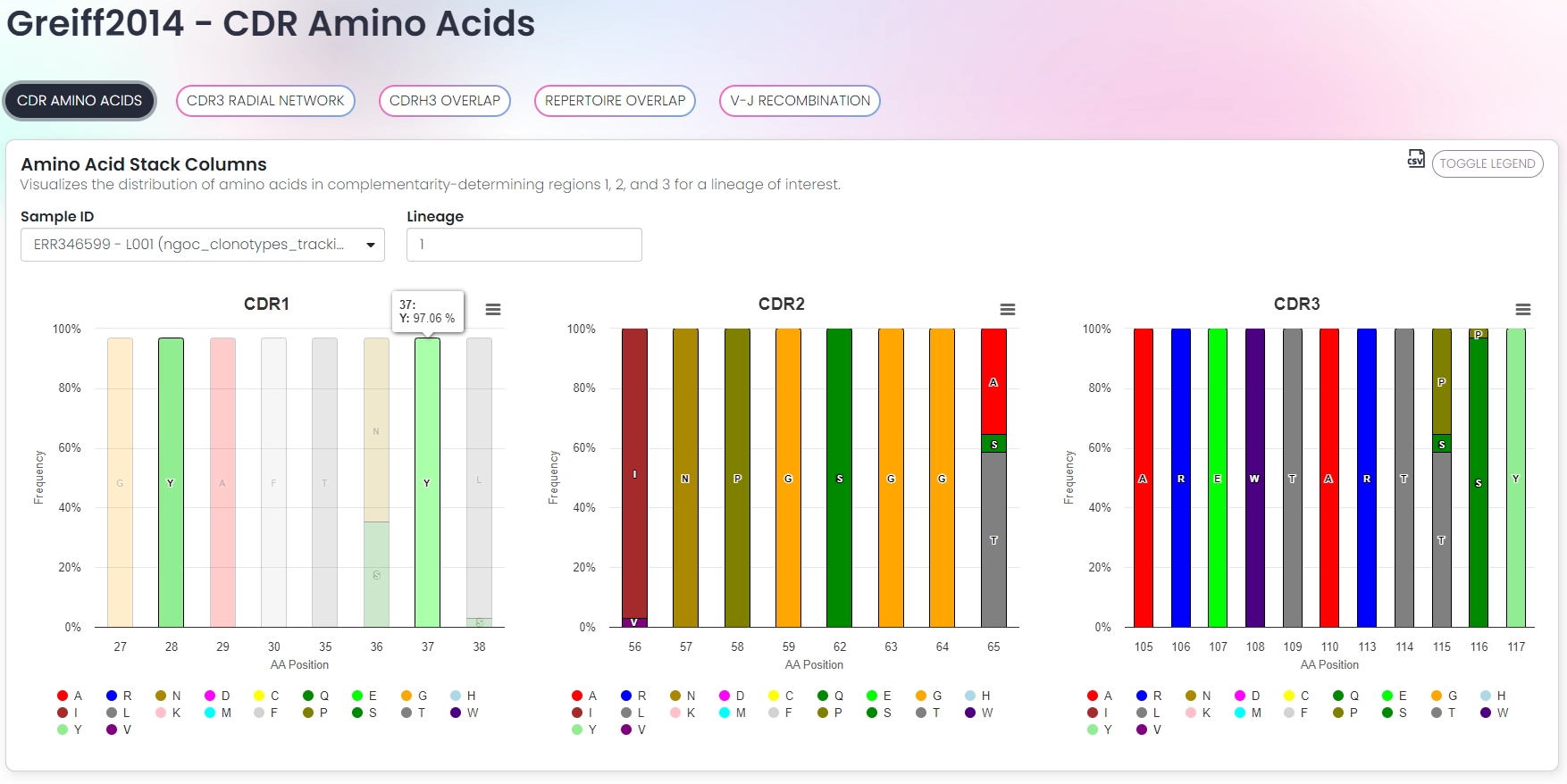
Interactive visualizations
Support for all common protocols (Illumina, UMI, 10x)
High-throughput data processing
Data security
Standardized according to AIRR community guidelines
ImmunoRaptor features
- Support for all common input data types such as BCR/TCR data from Illumina or 10X
- Assignment of clones/clonotypes/lineages
- Reconstruction of phylogenetic trees
- Analysis of CDR properties (e.g. amino acids, lengths, physicochemical properties)
- Analysis of somatic hypermutation rates
- Analysis of gene usage and V-J recombination
- Excelra global support
Start analyzing your immune repertoire sequencing data efficiently!